Bayesian model selection#
Author: Zeel B Patel
http://mlg.eng.cam.ac.uk/teaching/4f13/1920/bayesian finite regression.pdf (Last slide) http://mlg.eng.cam.ac.uk/teaching/4f13/1920/marginal likelihood.pdf
import numpy as np
import scipy.stats
import matplotlib.pyplot as plt
from matplotlib import rc
rc('font', size=14)
from scipy.optimize import minimize
import warnings
warnings.filterwarnings('ignore')
from time import time
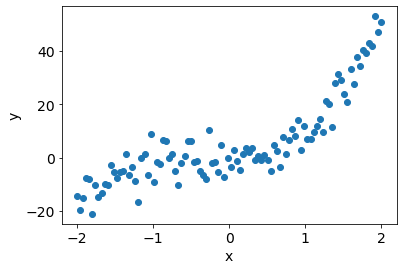
Pseudo-random data (degree 3 polynomial)#
np.random.seed(0)
N = 100
sigma_n = 5 # noise in data
sigma_w = 100 # parameter variance
x = np.linspace(-2,2,N).reshape(-1,1)
f_x = 4*x**3 + 3*x**2 + 2*x + 1
epsilon = np.random.normal(loc=0, scale=sigma_n, size=N).reshape(-1,1)
y = f_x + epsilon
plt.scatter(x, y);
plt.xlabel('x');plt.ylabel('y');
Marginal likelihood pdf#
def LogMarginalLikelihoodPdf(x, y):
return np.log(scipy.stats.multivariate_normal.pdf(y.squeeze(), np.zeros(N), (x@x.T)*sigma_w**2 + np.eye(N)*sigma_n**2))
Models#
x_M0 = x.copy()
x_M1 = np.hstack([np.ones((N,1)), x])
x_M2 = np.hstack([np.ones((N,1)), x, x**2])
x_M3 = np.hstack([np.ones((N,1)), x, x**2, x**3])
x_M4 = np.hstack([np.ones((N,1)), x, x**2, x**3, x**4])
x_M5 = np.hstack([np.ones((N,1)), x, x**2, x**3, x**4, x**5])
x_M6 = np.hstack([np.ones((N,1)), x, x**2, x**3, x**4, x**5, x**6])
x_M7 = np.hstack([np.ones((N,1)), x, x**2, x**3, x**4, x**5, x**6, x**7])
Model selection#
scores = [LogMarginalLikelihoodPdf(x_M, y) for x_M in [x_M0, x_M1, x_M2, x_M3, x_M4, x_M5, x_M6, x_M7]]
plt.plot(scores);
plt.xlabel('Degree of polynomial');
plt.ylabel('Log Marginal Likelihood \n(Higher is better)');
plt.xticks(range(len(scores)));
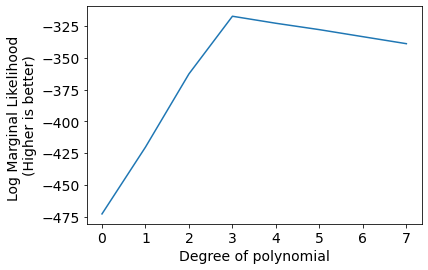
We can infer that Degree 3 polynomial is best suited to model current data.
Model selection with parameter optimization#
def NegLogMarginalLikelihoodPdf(params, x, y): # Negative log marginal likelihood (written in GP fashion)
sigma_n, sigma_w = params
K = (x@x.T)*sigma_w**2 + np.eye(N)*sigma_n**2
K_inv = np.linalg.pinv(K)
nll = 0.5*y.T@K_inv@y + 0.5*np.log(np.linalg.det(K)) + (len(y)/2)*np.log(2*np.pi)
return nll[0,0]
Negscores = []
sig_n_list = []
sig_w_list = []
sig_w = 10
sig_n = 10
for i, x_M in enumerate([x_M0, x_M1, x_M2, x_M3, x_M4, x_M5, x_M6, x_M7]):
init = time()
result = minimize(fun=NegLogMarginalLikelihoodPdf, x0=(sig_n, sig_w), args=(x_M, y))
sig_n_opt, sig_w_opt = result.x
sig_n_list.append(sig_n_opt)
sig_w_list.append(sig_w_opt)
print(f'degree = {i}, sigma_w={sig_w_opt}, sig_n_opt={sig_n_opt}', 'time:',time()-init,'seconds')
Negscores.append(NegLogMarginalLikelihoodPdf((sig_n_opt, sig_w_opt), x_M, y))
plt.plot(Negscores);
plt.xlabel('Degree of polynomial');
plt.ylabel('Neg Log Marginal Likelihood \n(Lower is better)');
plt.xticks(range(len(Negscores)));
degree = 0, sigma_w=11.386419730789676, sig_n_opt=10.409619988275981 time: 0.16215753555297852 seconds
degree = 1, sigma_w=8.895627576152828, sig_n_opt=8.940430356187846 time: 0.08962178230285645 seconds
degree = 2, sigma_w=7.119736606266272, sig_n_opt=6.9430152775634495 time: 0.14646410942077637 seconds
degree = 3, sigma_w=3.219312493175096, sig_n_opt=4.710523287770871 time: 0.14163684844970703 seconds
degree = 4, sigma_w=2.6041389205399996, sig_n_opt=4.739086634960693 time: 0.2660367488861084 seconds
degree = 5, sigma_w=-2.5131468222243427, sig_n_opt=4.745522282327305 time: 0.17346620559692383 seconds
degree = 6, sigma_w=2.163988485988779, sig_n_opt=4.768321344889457 time: 0.5917925834655762 seconds
degree = 7, sigma_w=1.6084504892174767, sig_n_opt=4.758818020810231 time: 0.32204294204711914 seconds
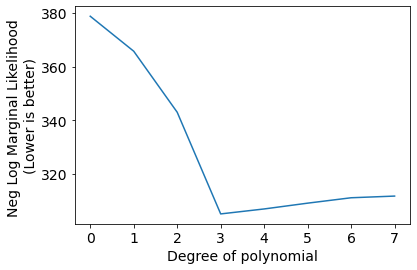
We can see the best set of hyper-parameters selected by gradient descent on negative log likelihood for indivisual models (degree of polynomial).
Drawbacks#
This method does not scale well with big data (while doing parameter optimization), because of matrix inversion.
Potential solution: optimize parameters with chepaer methods, then use Marginal likelihood for model selection
Using empirical bayes (Bishop)#
http://krasserm.github.io/2019/02/23/bayesian-linear-regression/
alpha = 1/sigma_w**2
beta = 1/sigma_n**2
Marginal likelihood pdf#
def posterior(Phi, t, alpha, beta, return_inverse=False):
"""Computes mean and covariance matrix of the posterior distribution."""
S_N_inv = alpha * np.eye(Phi.shape[1]) + beta * Phi.T.dot(Phi)
S_N = np.linalg.inv(S_N_inv)
m_N = beta * S_N.dot(Phi.T).dot(t)
if return_inverse:
return m_N, S_N, S_N_inv
else:
return m_N, S_N
def log_marginal_likelihood(Phi, t, alpha, beta):
"""Computes the log of the marginal likelihood."""
N, M = Phi.shape
m_N, _, S_N_inv = posterior(Phi, t, alpha, beta, return_inverse=True)
E_D = beta * np.sum((t - Phi.dot(m_N)) ** 2)
E_W = alpha * np.sum(m_N ** 2)
score = M * np.log(alpha) + \
N * np.log(beta) - \
E_D - \
E_W - \
np.log(np.linalg.det(S_N_inv)) - \
N * np.log(2 * np.pi)
return 0.5 * score
Bscores = [log_marginal_likelihood(x_M, y, alpha, beta) for x_M in [x_M0, x_M1, x_M2, x_M3, x_M4, x_M5, x_M6, x_M7]]
plt.plot(Bscores);
plt.xlabel('Degree of polynomial');
plt.ylabel('Log Marginal Likelihood \n(Higher is better)');
plt.xticks(range(len(Bscores)));

Results are exactly the same as the previous method.
Paramater tuning#
def fit(Phi, t, alpha_0=1e-5, beta_0=1e-5, max_iter=200, rtol=1e-5, verbose=False):
"""
Jointly infers the posterior sufficient statistics and optimal values
for alpha and beta by maximizing the log marginal likelihood.
Args:
Phi: Design matrix (N x M).
t: Target value array (N x 1).
alpha_0: Initial value for alpha.
beta_0: Initial value for beta.
max_iter: Maximum number of iterations.
rtol: Convergence criterion.
Returns:
alpha, beta, posterior mean, posterior covariance.
"""
N, M = Phi.shape
eigenvalues_0 = np.linalg.eigvalsh(Phi.T.dot(Phi))
beta = beta_0
alpha = alpha_0
for i in range(max_iter):
beta_prev = beta
alpha_prev = alpha
eigenvalues = eigenvalues_0 * beta
m_N, S_N, S_N_inv = posterior(Phi, t, alpha, beta, return_inverse=True)
gamma = np.sum(eigenvalues / (eigenvalues + alpha))
alpha = gamma / np.sum(m_N ** 2)
beta_inv = 1 / (N - gamma) * np.sum((t - Phi.dot(m_N)) ** 2)
beta = 1 / beta_inv
if np.isclose(alpha_prev, alpha, rtol=rtol) and np.isclose(beta_prev, beta, rtol=rtol):
if verbose:
print(f'Convergence after {i + 1} iterations.')
return alpha, beta, m_N, S_N
if verbose:
print(f'Stopped after {max_iter} iterations.')
return alpha, beta, m_N, S_N
for i, x_M in enumerate([x_M0, x_M1, x_M2, x_M3, x_M4, x_M5, x_M6, x_M7]):
init = time()
alpha, beta, m_N, S_N = fit(x_M, y, rtol=1e-5, verbose=True)
print('Degree',i,'sigma_w',1/np.sqrt(alpha),'sigma_n', 1/np.sqrt(beta))
print('Degree',i,'sigma_w',sig_w_list[i],'sigma_n', sig_n_list[i], '(Earlier method)')
print('time:',time()-init,'seconds')
Convergence after 3 iterations.
Degree 0 sigma_w 11.386046324847063 sigma_n 10.409625056741909
Degree 0 sigma_w 11.386419730789676 sigma_n 10.409619988275981 (Earlier method)
time: 0.0020685195922851562 seconds
Convergence after 3 iterations.
Degree 1 sigma_w 8.89562357924776 sigma_n 8.940424942304906
Degree 1 sigma_w 8.895627576152828 sigma_n 8.940430356187846 (Earlier method)
time: 0.006841897964477539 seconds
Convergence after 3 iterations.
Degree 2 sigma_w 7.119729723394079 sigma_n 6.943013815169844
Degree 2 sigma_w 7.119736606266272 sigma_n 6.9430152775634495 (Earlier method)
time: 0.0012140274047851562 seconds
Convergence after 4 iterations.
Degree 3 sigma_w 3.219311433389237 sigma_n 4.710523173354075
Degree 3 sigma_w 3.219312493175096 sigma_n 4.710523287770871 (Earlier method)
time: 0.0022554397583007812 seconds
Convergence after 6 iterations.
Degree 4 sigma_w 2.6041361190065677 sigma_n 4.739086719922639
Degree 4 sigma_w 2.6041389205399996 sigma_n 4.739086634960693 (Earlier method)
time: 0.0025718212127685547 seconds
Convergence after 9 iterations.
Degree 5 sigma_w 2.5131492993598394 sigma_n 4.745518078421809
Degree 5 sigma_w -2.5131468222243427 sigma_n 4.745522282327305 (Earlier method)
time: 0.003193378448486328 seconds
Convergence after 9 iterations.
Degree 6 sigma_w 2.1639963673023725 sigma_n 4.768325837603502
Degree 6 sigma_w 2.163988485988779 sigma_n 4.768321344889457 (Earlier method)
time: 0.0029621124267578125 seconds
Convergence after 6 iterations.
Degree 7 sigma_w 1.6084472283522588 sigma_n 4.758819295283589
Degree 7 sigma_w 1.6084504892174767 sigma_n 4.758818020810231 (Earlier method)
time: 0.002556324005126953 seconds
This method is extremely fast than the previous method. We get close answers while using any of the two methods.

